We believe in serving you, advancing health, and paving the way for great discoveries in biomedical research at the University of Michigan and beyond.
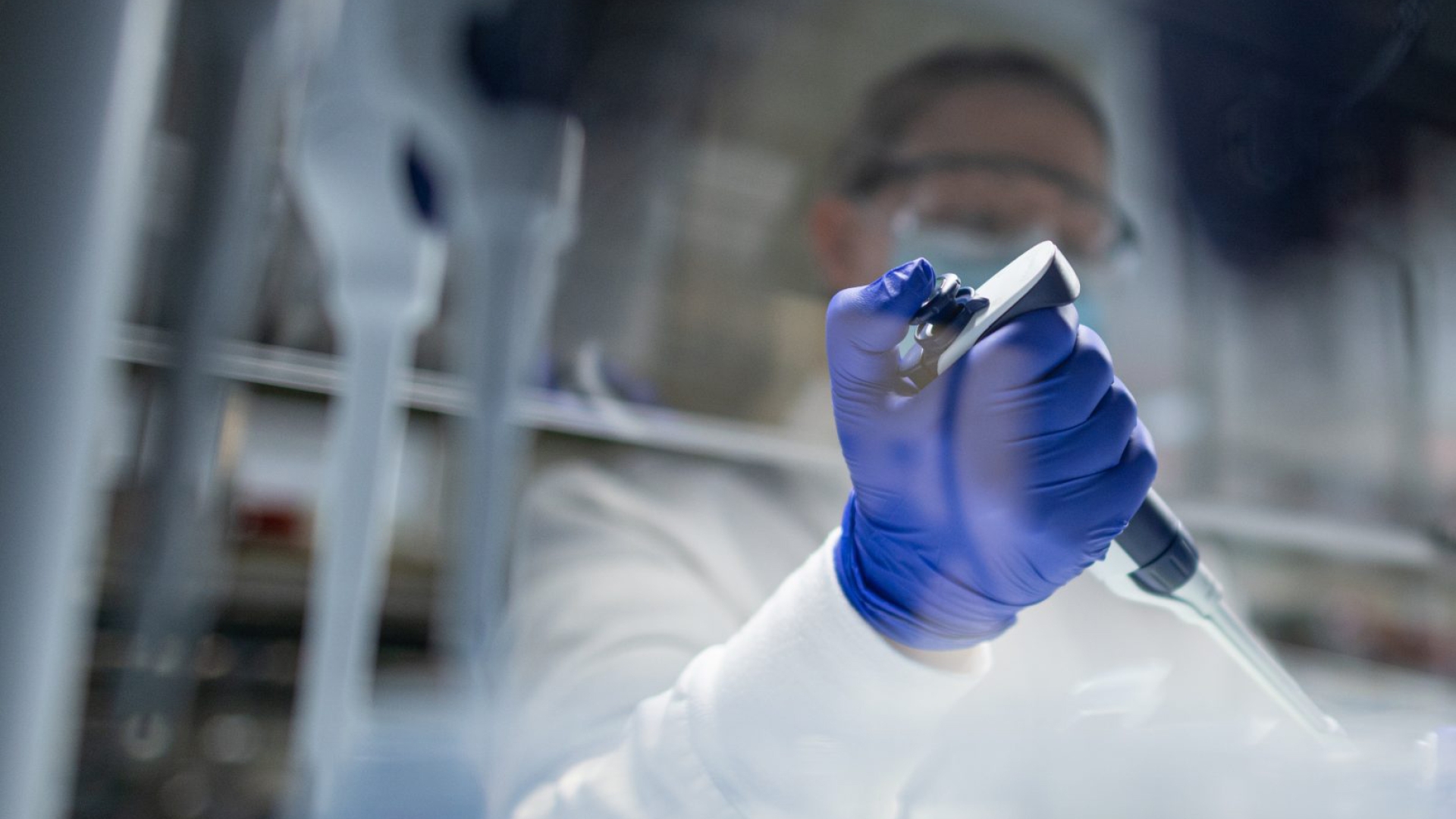

Cores are centralized facilities or labs which offer shared services, shared equipment, resources and expertise to biomedical researchers and investigators on a fee-for-service basis.
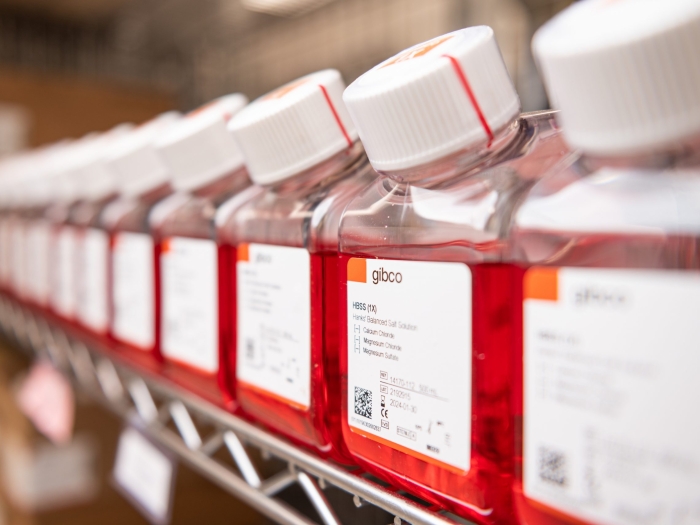

The Store provides University of Michigan research investigators with easy, on-site procurement of enzymes, reagents, and kits used in molecular, cell biology, and some protein chemistry.
More than 700 different items from 13 vendors are stocked at five locations for immediate purchase.
The BRCF centralized access to research services, shared equipment, and the expertise of biomedical researchers/investigators to support research projects. Explore our Careers, FAQs, and Mission here.
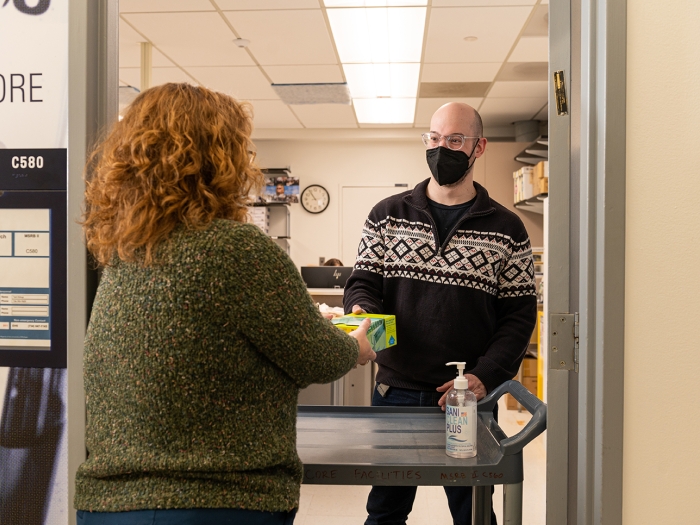

Several of the Cores in the Biomedical Research Core Facilities offers products for sale to help you with your research projects. From enzyme kits and reagents to mouse models and gene panels.
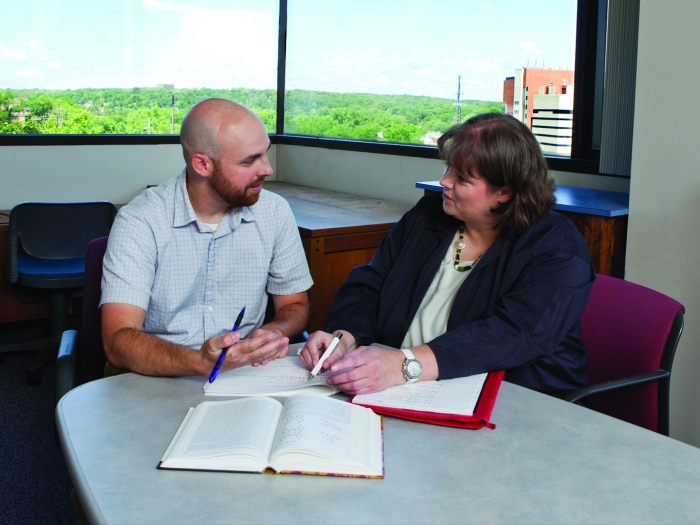

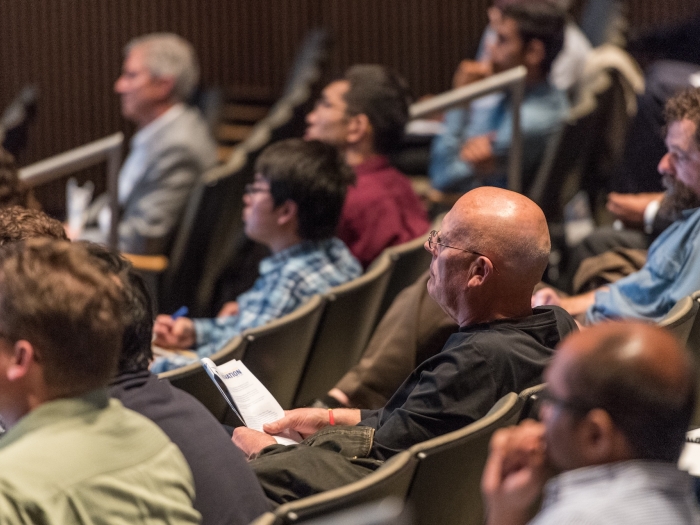
The BRCF can help you discover technology, find experts, and advance your research every step of the way. Consult with the Cores or attend a training event to collaborate with top experts and take advantage of the latest technology in research.
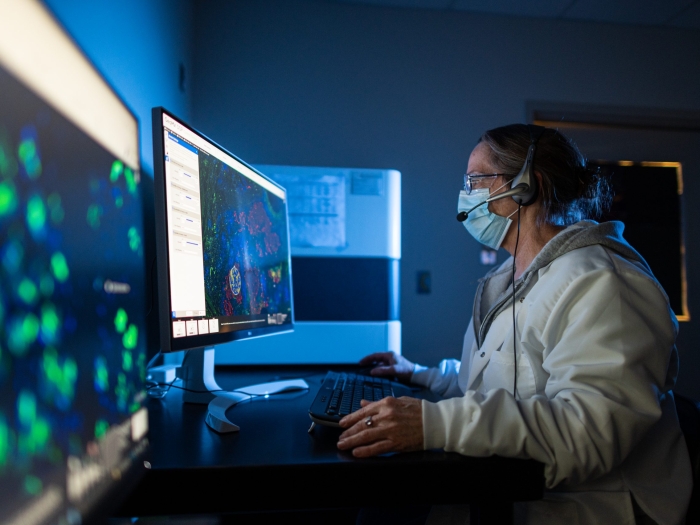

Whether you need to drop off a sample of DNA or cells for imaging, obtain expert analysis, or store your samples, the BRCF are here to assist you.
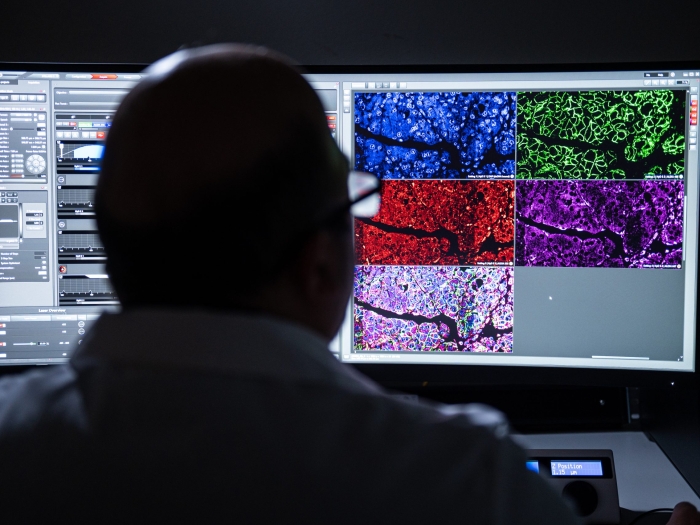

Cutting-edge technology and dozens of instruments in the fields of Flow Cytometry, and Microscopy Image & Analysis are available through the BRCF, and self-service and technician-operated options are available.
Purchase Research Materials
Several of the Cores in the Biomedical Research Core Facilities offers products for sale to help you with your research projects. From enzyme kits and reagents to mouse models and gene panels. For more information, please visit:
Consultation
The Biomedical Research Core Facilities (BRCF) can help you discover technology, find experts and advance your research every step of the way. Consult with any of our Cores to collaborate with top experts and take advantage of the latest technology in research. We help establish the scope and budget of your project, coordinate applications for funding, offer comprehensive training for investigators and more.
Many of our Cores offer their services to other academic and non-academic research institutions. Visit each Core’s specific website to learn more.
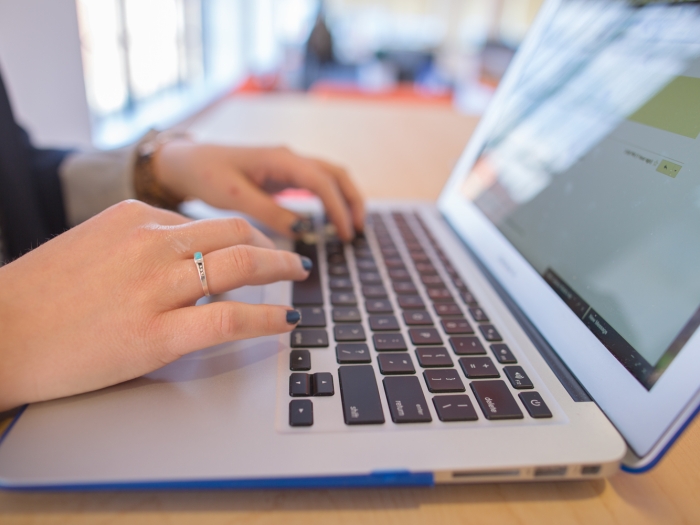
Prepare and Submit Your Samples & Data
Whether you need to drop off a sample of DNA or cells for imaging, the Biomedical Research Core Facilities are available to assist with your samples and data. Visit a specific core facility for more information.

Biomedical Research Core Facilities (BRCF), part of the University of Michigan Medical School Office of Research, are campus-wide laboratories that develop and provide state-of-the-art scientific resources to enable biomedical research. Our mission is that we believe in serving you, advancing health, and paving the way for great discoveries in biomedical research at the University of Michigan and beyond.
Due to the diversity of our work, many of our Cores have developed their own frequently asked questions (FAQs). Please visit each of our core websites to learn more about services and troubleshooting.
Because of the variety in services each Core provides, they all have a slightly different process to begin work with them. However, most project requests begin with a consultation. Check out the page of your core of interest to find out more, or view their “Contact Us” page to find out who to get in touch with.
Fill out our contact form and submit your question using “I have a project that may involve multiple cores.”
There are currently 11 cores under the BRCF umbrella, however many cores exist at the University of Michigan. To learn more about all of the cores located at U-M, please visit the Michigan Research Cores website.
MiCores is an operational software powered by Agilent Technologies and used by the BRCF to perform ordering, billing, and scheduling of scientific products and services. For more information about MiCores and how to register, visit the MiCores website.
There are many funding opportunities across the University and beyond. Highlighted below are some that are most applicable to the Biomedical Research Core Facilities. Please visit the UMMS Office of Research Funding Opportunities page for information about internal funding, external funding opportunities, bridging support, and more.
- Competition Space streamlines the process of finding and applying for, funding opportunities at the Medical School
- MICHR Funding: grant programs for basic, clinical, translational research and more available through the Michigan Institute for Clinical and Health Research (MICHR)
- Taubman Health Sciences Library Research Funding and Grants Guide
The Association of Biomolecular Resource Facilities (ABRF) has established recommended authorship guidelines that the Biomedical Research Core Facilities follow. Please see below for their recommended guidelines as implemented by the BRCF:
Why cite the contributions of a core?
It is important to document the use of each Biomedical Research Core Facility by investigators in publications. Doing so facilitates efforts to obtain funding for our Cores. Core facility personnel are valued scientists; when they make a substantial contribution to a publication, they deserve to be acknowledged like any other co-author. The existence of the BRCF relies on proper acknowledgment in publications and awards. Proper acknowledgment of the BRCF helps us to obtain support so that we may continue to provide services to the best of our abilities.
Please include the following information in the acknowledgments section of any publication resulting from services provided by or facilities used by any of the BRCF Cores:
- The name of the specific Biomedical Research Core Facility(ies) used in the scope of your research or project.
- Clearly, state how the author and core contributed to your research.
- If a center, department, grant or institution supported access to resources through funding or other means, please include that.
Statement Example
“In collaboration with this research, we acknowledge support from the University of Michigan Biomedical Research Core Facilities (Insert BRCF Core Name Here).”
If you have any questions about correctly attributing the Cores in a publication or award, please contact us.
For more information, please see the Authorship Guidelines established by the ABRF.
The BRCF administration office is located on the C-floor (C570) of Medical Science Research Building II (MSRB II). Each of our Cores is spread out across campus, including MSRB II, Biomedical Research Science Building (BSRB), Life Sciences Institute (LSI), Brehm, and North Campus Research Complex (NCRC).
How do I get to the MSRB II C-Floor?
The C-Floor can only be accessed by using the elevators in Medical Science Research Building II. Use the directions or map below to navigate to MSRB II.
Entering from 1150 W. Medical Center Dr. (MSRB II)
If you come through the main entrance at the circle drive, take the door to your left that says “MSRB II” and use those elevators to arrive at the C-Floor. You can also take the open staircase to the left of the entrance. View these directions on a map.
Entering from 1137 Catherine St. (Med Sci II)
If entering through the 4th floor of Medical Science Building II (Med Sci II), head left from the entrance and follow the hallway straight down until it dead-ends at a stairwell. Once through that doorway, take the small flight of stairs to your left onto the 4th floor of MSRB I. From there, follow the hallway to the right until you reach the MSRB II elevators. View these directions on a map. Similarly, from the main entrance to the 3rd floor of Medical Science Building II (Med Sci II), head right down the long hallway until it dead-ends at a stairwell. Once through that doorway, take the small flight of stairs to your left onto the 3rd-floor MSRB I. From there, follow the hallway to the right until you reach the MSRB II elevators. View these directions on a map.
Entering from the Cancer Center of University Hospital
If entering from the Cancer Center parking structure, take the elevators to the 7th-floor skywalk and take a right at the first set of doors (this is the 3rd floor of Med Sci I). This will take you down a long hallway that opens up into Med Sci II. Take a right at the vending machines until you reach the stairwell. Once through that doorway, take the small flight of stairs to your left onto the 3rd floor of MSRB I. From there, follow the hallway to the right until you reach the MSRB II elevators. Follow these same directions if coming directly from the Cancer Center or UH. You must be on the 2nd floor of either building to connect to the 3rd floor of Med Sci I and II. View a more detailed map indicating directions from multiple sites at Michigan Medicine (note, this is a large file).

Ann Arbor, MI 48109-0674
Subscribe today to receive the latest news, core updates, monthly newsletter, and more from the BRCF.





At Michigan Medicine, we unequivocally recognize racism as a public health issue and we stand together against bias and inequality.