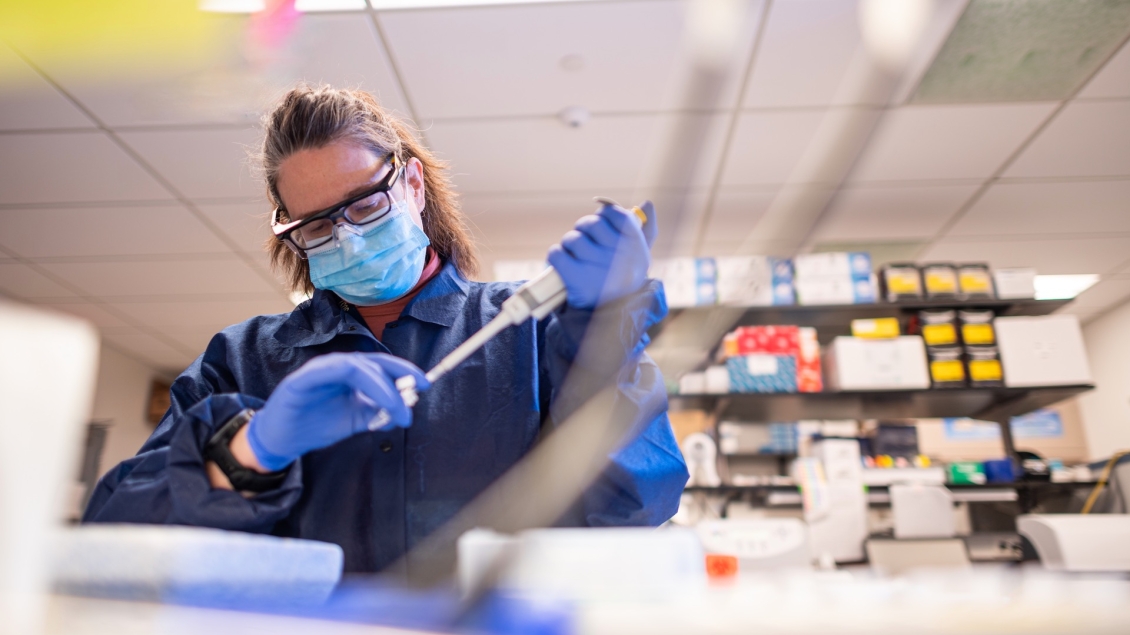
Preparing samples for analysis in epigenetic regulation in both genome-wide and locus-specific manners.
The Epigenomics Core provides resources and services to prepare samples for analysis in epigenetic regulation in both genome-wide and locus-specific manners. Epigenetic regulation refers to DNA sequence-independent regulation of heritable traits that impacts gene expression. In recent years the term has broadened to also encompass processes that control gene expression and genomic functions beyond DNA sequence but which may be more transient or facultative in nature.

providing information on the 3D organization of the genome and its role in gene regulation

providing information on DNA methylation status across the genome at different levels of resolution
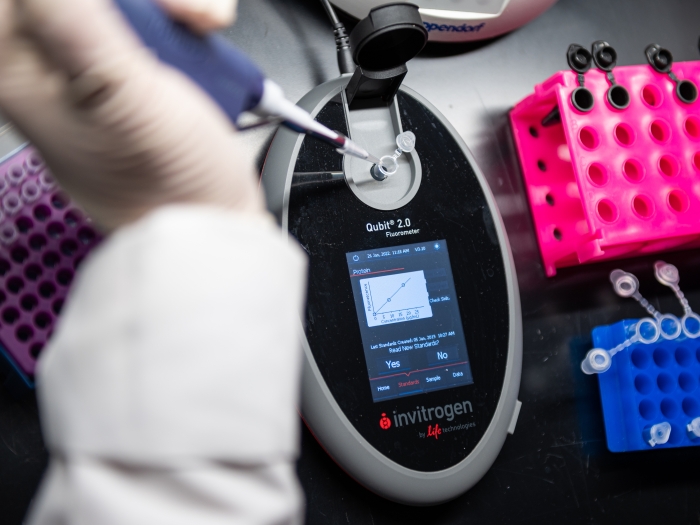

mapping genome-wide distribution patterns of specific histone modifications

identifying accessible regions in the chromatin structure
| Service Requested | Unit | Internal Rate (U-M |
| Sample Quality Control only (Qubit and TapeStation) | per sample | $28 |
| Sample Quality Control and Bisulfite Treatment | per sample | $34 |
| PCR Reaction (includes QC on TapeStation) | per sample | $29 |
| DNA Extraction & Quality Control | per sample | $40 |
| Cryopulverization of flash frozen tissues (materials + labor) | per sample | $52 |
| nuclei preparation from flash frozen tissues (includes cryopulverization material and labor) | per sample | $78 |
| nuclei preparation from fresh or cryopreserved cells (materials + labor) | per sample | $46 |
| Service Requested | Unit | Internal Rate (U-M) |
| WGBS (includes sample QC and library QC/quant) | per assay | $277 |
| mERRBS (includes sample QC and library QC/quant) | per assay | $267 |
| ChIP-seq library (includes sample QC and library QC/quant) | per assay | $212 |
| ATAC-Seq library prep (includes library QC/quant; nuclei prep depends on material) | per assay | $236 |
| Amplicon Bisulfite-Seq library preparation (library QC/quant) | per assay | $90 |
| 5hMeDIP-Seq (includes sample QC and library QC/quant) | per sample | $235 |
| Sample Preparation Labor Rate | per hour | $121 |
| Pyrosequencing | full plate | $363 |
| OxBS-Seq Preparation | per sample | contact us |
Please Note: Sequencing costs are based on Advanced Genomics Core rates.
| NovaSeq 6000 | Unit | Internal Rate (U-M) |
| Illumina NovaSeq 6000 (SP) 100 Cycle Sequencing | per flow cell | $3,972 |
| Illumina NovaSeq 6000 (SP) 300 Cycle Sequencing | per flow cell | $4,973 |
| Illumina NovaSeq 6000 (SP) 500 Cycle Sequencing | per flow cell | $6,309 |
| Illumina NovaSeq 6000 (S1) 100 Cycle Sequencing | per flow cell | $6,292 |
| Illumina NovaSeq 6000 (S1) 200 Cycle Sequencing | per flow cell | $7,405 |
| Illumina NovaSeq 6000 (S1) 300 Cycle Sequencing | per flow cell | $7,850 |
| Illumina NovaSeq 6000 (S2) 100 Cycle Sequencing | per flow cell | $10,471 |
| Illumina NovaSeq 6000 (S2) 200 Cycle Sequencing | per flow cell | $12,418 |
| Illumina NovaSeq 6000 (S2) 300 Cycle Sequencing | per flow cell | $13,086 |
| Illumina NovaSeq 6000 (S4) 200 Cycle Sequencing | per flow cell | $18,476 |
| Illumina NovaSeq 6000 (S4) 300 Cycle Sequencing | per flow cell | $20,408 |
One array requires 16 samples. Cost is greater if fewer than 16 hybridizations are requested. For pricing please contact [email protected].
Included in the service request to the Epigenomics Core is the following output:
- Raw sequence data (FASTQ files)
- FASTCQ output (HTML and TXT files)
For ERRBS/mERRBS and WGBS projects the following files are also provided:
- CpG context methylation calls (TXT files)
- CpG context methylation calls for visualization (BEDGRAPH files)
- Summary statistics (Excel file)
For a more in-depth analysis, or if you require custom output, please state your interest at the consultation.
| Authors | Year | Publication |
| Straight, B, Fisher, G, Needham, BL, Naugle A, Olungah C, Wanitjirattikal P, Root C, Farman J, Barkman T, Lalancette C. | 2019 | Lifetime stress and war exposure timing may predict methylation changes at NR3C1 based on a pilot study in a warrior cohort in a small‐scale society in Kenya. Am J Hum Biol. 2020; e23515. |
| Huber AK, Patel N, Pagani CA, Marini S, Padmanabhan KR, Matera DL, Said M, Hwang C, Hsu GC-Y, Poli AA, Strong AL, Visser ND, Greenstein JA, Nelson R, Li S, Longaker MT, Tang Y, Weiss SJ, Baker BM, James AW, Levi B. | 2019 | Immobilization after injury alters extracellular matrix and stem cell fate. Published October 1, 2020; First published July 16, 2020, J Clin Invest. 2020; 130(10): 5444-5460. |
Acknowledgment of Contributions
Projects that use data or samples generated by the Epigenomics Core and result in publication should include an acknowledgement of the Core or include members as coauthors. The Epigenomics Core uses these acknowledgements and authorship to help demonstrate our contributions to the research community. This in turn helps secure future funding to maintain a robust Core facility and provide professional development of its staff. The Association of Biomolecular Resource Facilities has published a guideline to use when considering whether or not to include core laboratory members on publications.


1150 West Medical Center Drive
Ann Arbor, MI 48109